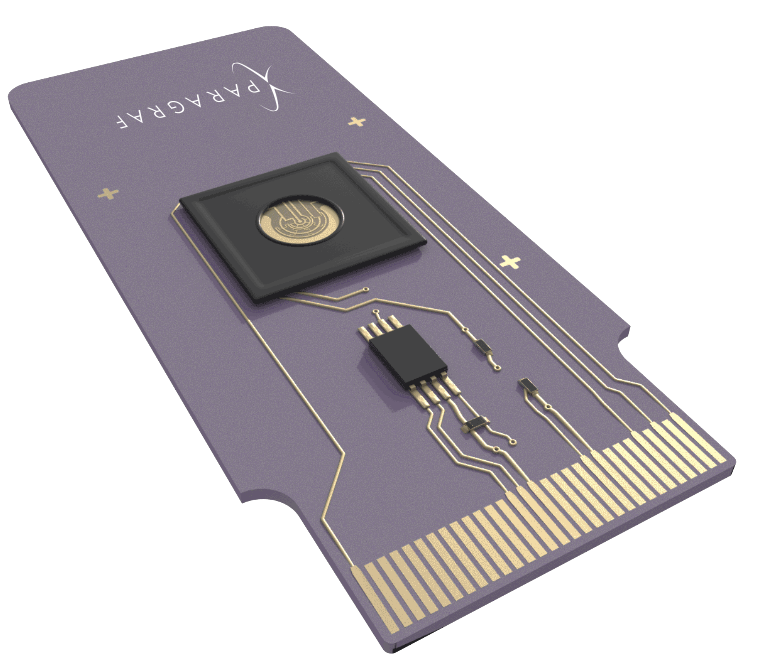
The BPU Platform consists of:

What is a BPU?
A BPU is an electrical chip that is a type of microprocessor. It does not process digital signals. Instead, it processes biological signals or “biosignals.” What this means is that a BPU converts biological interactions such as an antibody binding a virus into the kind of electrical signals we’re used to seeing in conventional electronics.
Biosignal Bandwidth
You may be familiar with the term “biomarker.” If a biomarker is a picture of the current state of some biology, then a biosignal is a video of that state of biology. It’s adding the real-time component of recording to our familiar understanding of biomarkers.
Some examples of interactions recorded in biosignals are binding of antibodies to viral proteins and strands of DNA binding to complementary DNA. However, there are a tremendously large number of potential biosignals that could be measured. A BPU can be characterized by it’s biosignal bandwidth. The biosignal bandwidth is roughly the number of types of biological interactions that can be monitored on one chip, multiplied by the number of input or output modes available to each of those interactions.
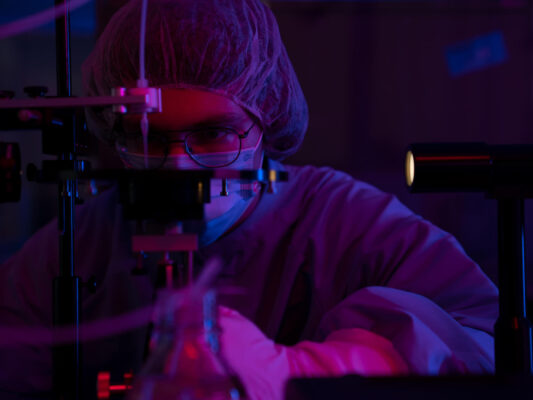
Real-time Multiomics
As biological research has synthesized genomics, proteomics, metabolomics and transcriptomics into systems biology, a new multiomics approach to biological research has emerged. Today, multiomics studies are challenging and expensive.
Paragraf has developed a platform for experimental work that unifies the multiple omics approaches to measurement.
This increases access to multiomics data by enabling more individual labs to successfully attempt multiomics studies.
As we are building semiconductor-based BPUs, we can package the much-smaller transistors later so the BPU can work off the same sample; but for now we just run more BPUs next to each other. Our platform allows us to measure DNA, RNA and proteins as the only platform in production. And we do already have next-gen BPUs with multiplexing – more measurements next to each other – in small-batch production for our Innovation R&D groups.
Today, we run one real-time multiomics measurement a lot and that is RNA binding proteins (e.g., gRNA to CAS protein binding).

Paragraf’s BPU is the only tech that can real-time measure this central and important type of multiomics that otherwise is considered impossible to measure with current state technology.
Almost 20 years ago, the Human Genome Project verified something that greatly complicated the way we understand biology: that proteins are not derived from simple expression of the genetic codes in nucleic acid sequences. The human genome contains open reading frames that code for roughly 20,000 canonical proteins. Around 70,000 additional proteins can be created by using alternative splicing of the DNA code. However, to our best understanding, a single human cell contains at least 1,000,000 different forms of proteins.
The elegance of the central dogma of biology is that information can be transferred from DNA to RNA to protein directly, yet it is now augmented by an understanding that reality is much more complicated. The life cycle of a particular protein is constructed by overlapping parameters such as gene regulation; post-transcriptional and post-translational modifications, which are regulated by interactions with transcription factors; enzymes; carrier proteins and other nucleic acids. In addition, the function of the same protein requires a multitude of interactions between other proteins, small molecules and metabolic markers. Complexity in biology comes not only from the complexity of the networks of interactions involved but also in the variations in the parts of those networks. This compounded complexity creates limits to what can be accomplished via reductionist approaches to biology, as well as limits to the usefulness of naïve mappings of engineering and computer science concepts into biology.
This understanding led to the creation of the field of systems biology, the study of emergent patterns from the dynamic complexity of biology rather than focus on a particular type of molecule, such as genomics, proteomics or metabolomics. An early editorial in Science put it well: “the pluralism of causes and effects in biological networks is better addressed by observing, through quantitative measures, multiple components simultaneously, and by rigorous data integration with mathematical models.” This powerful concept offers a way forward to understand how observable traits come about in biological systems and the creation of a new approach to measurement: multiomics.
The current approach to multiomics studies is rooted in a scale up of single omics biological techniques. In the recent years, collections of large multiomics datasets have enabled population studies that captures markers for disease diagnosis, lifestyle and environmental conditions to better inform individualized and precise medical treatments. One major factor is the challenge to integrate information from independent omics studies into actionable insights. The analysis of muliomics data requires complex mining of the relations among different biological process streams using maching learning methods and multiperspective analysis.

While tremendous strides in increasing throughput and reducing cost have enabled multiomics studies through collaborative research, integration of simultaneous measurements from multiple biological components is not currently possible. Platforms of the various omics techniques including genomics, proteomics, transcriptomics, metabolomics and cytomics are used in coordinated studies to produce snapshots of data with an intention of creating a holistic system level understanding. These studies can be quite difficult, normally requiring months or years to accomplish. There are outstanding difficulties in experiment planning, data integration and cost control when performing multiomics studies. Even with the heroic efforts of these studies, the dynamic nature of biology is not addressed by current approaches, and computer models are required to simulate dynamic responses between individual time points.
Multiomics
At the same time that multiomics was being developed as an experimental foundation for systems biology, nanoelectronics was being developed as a foundation to directly link digital electronics to active biological systems. The combination of a unifying multiomics analytical biosensor with machine learning algorithms would be a very effective tool for complex biological data collection and decisive interpretation. Of the various designs and materials demonstrated over the 20 years of this effort, graphene transistor-based sensors are uniquely positioned to provide the capability to detect multiple analytes via an integrative device, thereby enabling greater accessibility to an academic research lab on a wider commercial scale.
Apply to partner with us!
Our growing network of partners operate in markets across diagnostics, research, agriculture, defense and more – all leveraging Paragraf’s BPU Platform to build unrivalled products that create and disrupt their markets.
To partner with Paragraf, please let us know a little bit about you, your team, the idea you want to commercialize with us and what will make you our next great partner.


